Moving diagnostic frontiers in dermatology
The demand for precision diagnostics
Skin diseases impair patients’ quality of life more drastically than cancer and diabetes²’
Your point-of-need molecular solution
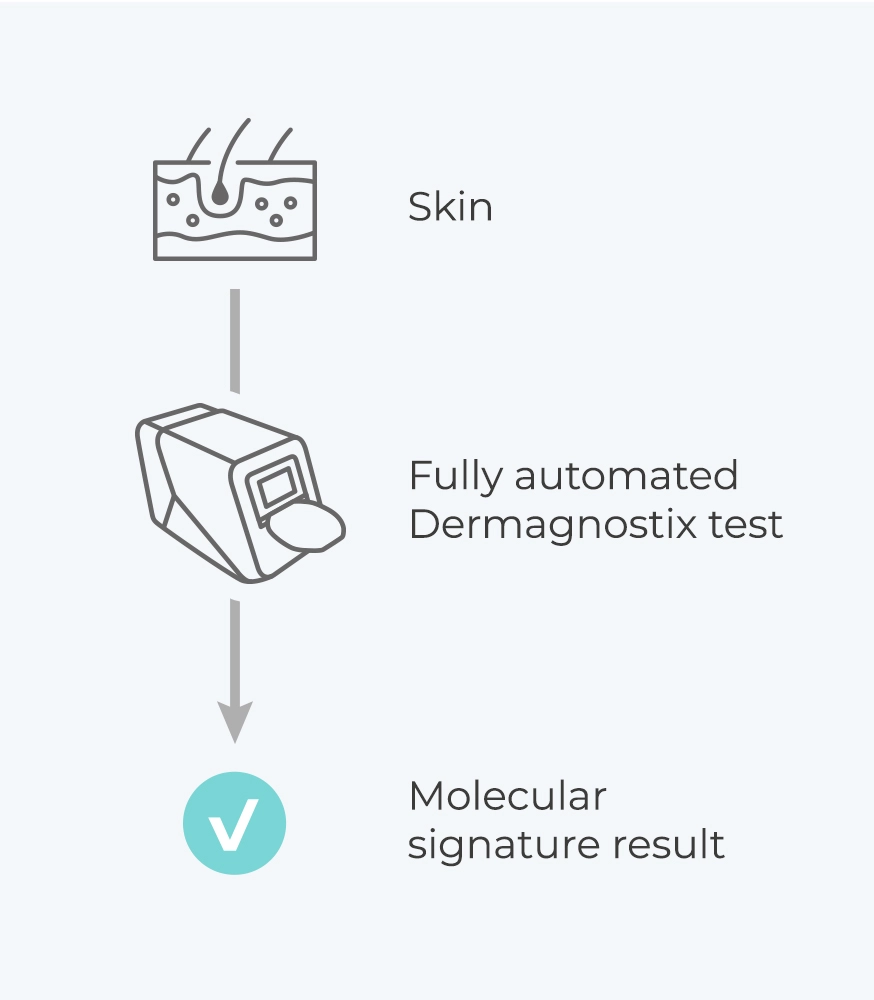
Groundbreaking translational research meets cutting-edge technology – Dermagnostix provides unique diagnostic tests for unmet clinical needs in dermatology on a fully automated portable laboratory. With PsorX (CE-IVD), Dermagnostix offers the first in-class molecular test for differential diagnosis of psoriasis and eczema.

Simple diagnostic tools for complex skin diseases
Simple
Ensuring maximum usability, the user only needs to place the sample into the LabDisk and insert the LabDisk into the Analyzer.
Quick
Results are delivered in only 45 min to 2 hours enabling physicians to make timely decisions about their patients’ therapies.
Cost-efficient
Each LabDisk contains only minimum amounts of reagents and materials due to microfluidics design.
Precise
All advantages of centrifugal microfluidics are exploited, enabling reliable, objective and valid results.
Point of need
Timely diagnoses where they are needed: Get objective and valid results anywhere and at any time.
Product of Germany
Our technology is developed, produced and tested in Germany.
Meet the Dermagnostix team
Get our news
Dermagnostix expands US business development by Eureka Innowwide funding and acceptance into German Accelerator
By participating in several conferences and trade shows in the US – most recently at the ISDP meeting in San Diego – Dermagnostix has already been able to show that the interest in PsorX is high and the feedback is consistently positive. In 2024, the focus will shift even more to business development in the USA. The
Upcoming Events
References
- Flohr C, Hay R (2021): Putting the burden of skin diseases on the global map. Br J Dermatol 184(2): 189-190.
- Montero-Vílchez T, Sánchez-Díaz M, Martínez-López A, Arias-Santiago S (2021): Quality of Life in Patients with Skin Disease and Their Cohabitants. Health-Related Quality of Life Measurement Tools, Predictors and Modifiers. IntechOpen.
- Augustin M, Glaeske G, Hagenström K (2021): Neurodermitisreport - Prävention, Versorgung und Innovation.
